Warning
This documents an unmaintained version of NetworkX. Please upgrade to a maintained version and see the current NetworkX documentation.
from_numpy_matrix¶
-
from_numpy_matrix(A, parallel_edges=False, create_using=None)[source]¶ Return a graph from numpy matrix.
The numpy matrix is interpreted as an adjacency matrix for the graph.
Parameters: - A (numpy matrix) – An adjacency matrix representation of a graph
- parallel_edges (Boolean) – If this is
True,create_usingis a multigraph, andAis an integer matrix, then entry (i, j) in the matrix is interpreted as the number of parallel edges joining vertices i and j in the graph. If it isFalse, then the entries in the adjacency matrix are interpreted as the weight of a single edge joining the vertices. - create_using (NetworkX graph) – Use specified graph for result. The default is Graph()
Notes
If
create_usingis an instance ofnetworkx.MultiGraphornetworkx.MultiDiGraph,parallel_edgesisTrue, and the entries ofAare of typeint, then this function returns a multigraph (of the same type ascreate_using) with parallel edges.If
create_usingis an undirected multigraph, then only the edges indicated by the upper triangle of the matrix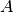 will be added to the
graph.
will be added to the
graph.If the numpy matrix has a single data type for each matrix entry it will be converted to an appropriate Python data type.
If the numpy matrix has a user-specified compound data type the names of the data fields will be used as attribute keys in the resulting NetworkX graph.
See also
Examples
Simple integer weights on edges:
>>> import numpy >>> A=numpy.matrix([[1, 1], [2, 1]]) >>> G=nx.from_numpy_matrix(A)
If
create_usingis a multigraph and the matrix has only integer entries, the entries will be interpreted as weighted edges joining the vertices (without creating parallel edges):>>> import numpy >>> A = numpy.matrix([[1, 1], [1, 2]]) >>> G = nx.from_numpy_matrix(A, create_using = nx.MultiGraph()) >>> G[1][1] {0: {'weight': 2}}
If
create_usingis a multigraph and the matrix has only integer entries butparallel_edgesisTrue, then the entries will be interpreted as the number of parallel edges joining those two vertices:>>> import numpy >>> A = numpy.matrix([[1, 1], [1, 2]]) >>> temp = nx.MultiGraph() >>> G = nx.from_numpy_matrix(A, parallel_edges = True, create_using = temp) >>> G[1][1] {0: {'weight': 1}, 1: {'weight': 1}}
User defined compound data type on edges:
>>> import numpy >>> dt = [('weight', float), ('cost', int)] >>> A = numpy.matrix([[(1.0, 2)]], dtype = dt) >>> G = nx.from_numpy_matrix(A) >>> G.edges() [(0, 0)] >>> G[0][0]['cost'] 2 >>> G[0][0]['weight'] 1.0